Lines of Code
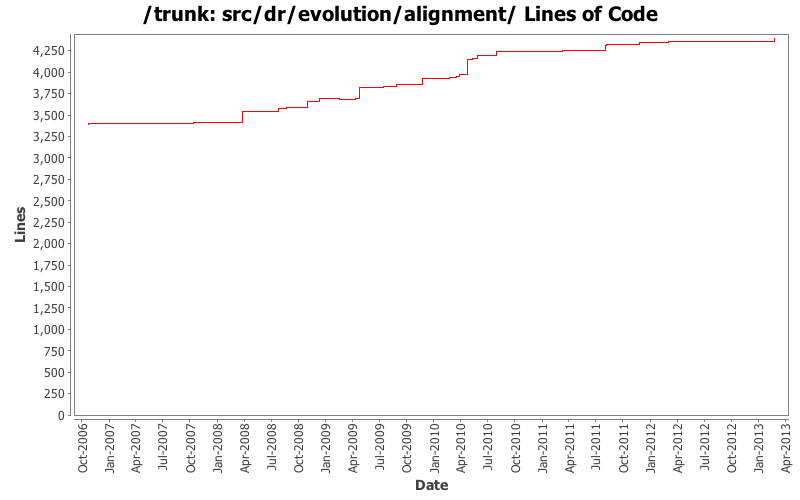
[root]/src/dr/evolution/alignment

| Author | Changes | Lines of Code | Lines per Change |
|---|---|---|---|
| Totals | 66 (100.0%) | 2445 (100.0%) | 37.0 |
| alexei.drummond | 7 (10.6%) | 1001 (40.9%) | 143.0 |
| rambaut | 30 (45.5%) | 976 (39.9%) | 32.5 |
| alexander.alekseyenko | 3 (4.5%) | 161 (6.6%) | 53.6 |
| msuchard | 10 (15.2%) | 116 (4.7%) | 11.6 |
| akaruiws | 1 (1.5%) | 50 (2.0%) | 50.0 |
| jheled | 4 (6.1%) | 45 (1.8%) | 11.2 |
| dong.w.xie@gmail.com | 2 (3.0%) | 39 (1.6%) | 19.5 |
| dong.w.xie | 2 (3.0%) | 36 (1.5%) | 18.0 |
| fbielejec | 6 (9.1%) | 17 (0.7%) | 2.8 |
| sibon.li | 1 (1.5%) | 4 (0.2%) | 4.0 |
trunk: merge Issue 677 : Using Microsatellite loci for inferring one phylogenetic tree that uses all the loci.
39 lines of code changed in 2 files:
Porting fixes from 1.7.3 back into the trunk
4 lines of code changed in 2 files:
Commented out all of my last commit until issues resolved
4 lines of code changed in 1 file:
setter for count report - safe
3 lines of code changed in 2 files:
setter for count report
1 lines of code changed in 1 file:
setter for count report
6 lines of code changed in 1 file:
trunk:solve Issue 457: BEAST should raise an error if binary dataType set but data is not binary, and Issue 312: BEAST continues to run (inappropriately) when sequence contains invalid character
27 lines of code changed in 1 file:
new constructor
7 lines of code changed in 2 files:
added handling for multi-block partitions
66 lines of code changed in 1 file:
seqgen generalization prep
11 lines of code changed in 1 file:
Hack to allow CompleteHistorySimulator to print out GeneralDataTypes. Several summary statistics in GeneralDataType still needs to be fixed when a sequence is a list of Strings and not chars.
4 lines of code changed in 1 file:
Classes for modelling the relative rates of loci.
50 lines of code changed in 1 file:
Fixed bug in ascertainment correction (miscounted) and implemented in BEAGLE. Genomic-wide SNPs here we come!
43 lines of code changed in 2 files:
Fixed an issue with root optimization for some trees in PathOGen.
6 lines of code changed in 1 file:
More work on the hypermutation model.
62 lines of code changed in 2 files:
Created an HypermutantAlignment that flags hypermutation contexts for use in the APOBECErrorModel. It does this by replacing 'A's by an ambiguity code for A/G if it is in the correct context in the alignment. These can then be compressed into unique site patterns so that speed is not affected.
Extracted a common abstract base class with ConvertAlignment which wraps an alignment.
477 lines of code changed in 4 files:
Patched ConvertAlignment so that it will convert a codon alignment into nucleotides.
24 lines of code changed in 1 file:
Can now simulate (and report) the complete history of any model in BEAST, most importantly codon models (with branch rate variation) and report the true number of synonymous and nonsynonymous mutations in a synthetic dataset.
9 lines of code changed in 1 file:
Fixed a small bug that prevented beagle from using multiple instances when certain patterns are used.
30 lines of code changed in 1 file:
Allow parser options to *not* condense site patterns into a unique list; this is necessary for robust counting of S and N mutations
9 lines of code changed in 1 file:
Tested alignment statistics; appear fine.
4 lines of code changed in 1 file:
Implemented more alignment stats in BasicAlignment.toString(). A site is considered informative if at least 2 states are observed at least 2 times. A site is considered a singleton if it is not conserved and not informative.
65 lines of code changed in 1 file:
BEAST now counts number of invariant sites
13 lines of code changed in 2 files:
automatic idea changes and generify
15 lines of code changed in 2 files:
estimating initial root height from node calibrations
288 lines of code changed in 1 file:
Update Big Change of BEAUTi: fix bugs in generator and change default values. But some of operators are not showing.
9 lines of code changed in 1 file:
Added option to SitePatternsParse that disables stripping out purely ambiguous patterns; these have some value when testing priors.
10 lines of code changed in 1 file:
Extended linking/unlinking to include trees by mirroring the setup we use for substitution models. Not complete yet so have included a static switch in DataPanel.ALLOW_UNLINKED_TREES which when false, reverts to the old GUI.
3 lines of code changed in 4 files:
Made some improvements to TaxonList and Taxa to make them work better with collections. For a start TaxonList is now Iterable so can be used in a for(Taxon taxon : taxonList) construct. In the long run we would be better dropping TaxonList and just replacing it with List<Taxon>.
154 lines of code changed in 10 files:
The "pattern of site" mapping is broken. I appriciate it if someone more familiar with this code audit my fix.
9 lines of code changed in 1 file:
Fixed problem with ConvertAlignment not using the correct genetic code (it basically always converted with the Universal code even if specified otherwise in the parser). This would result in incorrect likelihoods for other genetic codes in Codon models.
38 lines of code changed in 1 file:
automatic code improvments (intellij)
21 lines of code changed in 1 file:
Created a new parser called 'patternSubSet' which takes a SiteList (which includes Alignments and SitePatters) and creates a subset.
1 lines of code changed in 1 file:
Option added to patterns to divide the pattern list up into subsets for more efficient distribution amongst threads when not partitioned.
70 lines of code changed in 1 file:
Was finding that for pure R/Y encoded data the empirical frequency estimation was getting stuck in an iteration that never converged. Added a maximum number of iterations. Also added the MrBayes estimator (as opposed to the current one which matches PAUP) but this is not on by default.
82 lines of code changed in 2 files:
small bug fix in ascertained site patterns parser + RLTV improvements
53 lines of code changed in 1 file:
Painted a few more details of the new BEAUti. It now generates a BEAST XML from a Nexus file with charPartitions defined, but the BEAST XML doesn't run yet :-)
456 lines of code changed in 1 file:
Added classes necessary for MSSD a.k.a. Aquisition-Loss-Substitution (ALS) a.k.a. Immigration-Mutation-Death model functionality. Added prior and integrated immigration prior correction for alternative splicing creation and loss model.
AscertainedSitePatterns -- can handle ascertainment of two types
MutationDeathType -- extends a datatype with an additional state for the death state. Slightly differently (more clean?) structured than the other datatypes in the project. Let me know if you like it.
MutationDeathModel -- extends an arbitrary SubstitutionModel to include an additional absorbing death state.
ScaledTreeLengthRateModel -- Scales the sum of all edges to a specified value (like 1.0 in my case)
testBinary.xml -- a small example of who all of this stuff work
Changes:
BeastParser: added the parsers here
SitePatterns: made comparePatterns() protected instead of private
FrequencyModel: added a function to get the frequency parameter
131 lines of code changed in 2 files:
Moved NodeRateProvider and BranchRateProvider to dr.evolution.tree because dr.evolution.* should not depend on dr.evomodel.* (plus minor other tidying)
124 lines of code changed in 1 file:
Updated manuscript and manual and cleaned up a bit of lint...
3 lines of code changed in 1 file:
(1 more)