Lines of Code
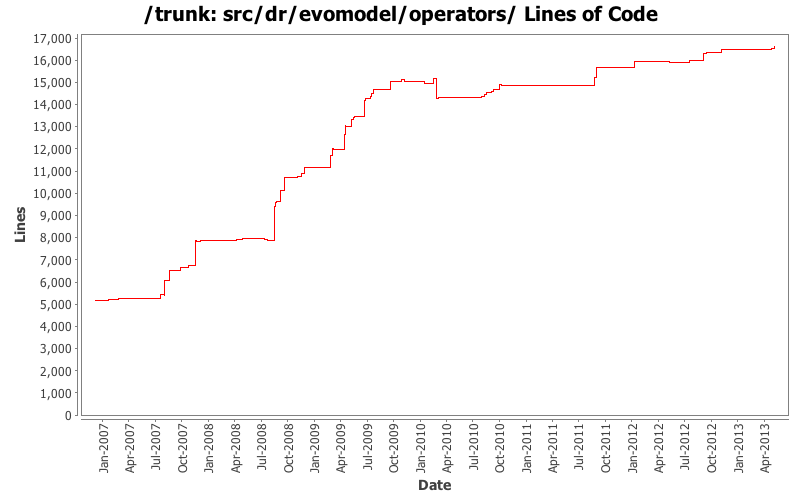
[root]/src/dr/evomodel/operators

| Author | Changes | Lines of Code | Lines per Change |
|---|---|---|---|
| Totals | 366 (100.0%) | 22411 (100.0%) | 61.2 |
| Sebastian.Hoehna | 46 (12.6%) | 6364 (28.4%) | 138.3 |
| alexei.drummond | 53 (14.5%) | 4146 (18.5%) | 78.2 |
| jheled | 60 (16.4%) | 3817 (17.0%) | 63.6 |
| msuchard | 65 (17.8%) | 3242 (14.5%) | 49.8 |
| rambaut | 68 (18.6%) | 1834 (8.2%) | 26.9 |
| graham853@gmail.com | 30 (8.2%) | 1755 (7.8%) | 58.5 |
| Michael.DefoinPLatel | 5 (1.4%) | 658 (2.9%) | 131.6 |
| dong.w.xie | 37 (10.1%) | 402 (1.8%) | 10.8 |
| aaron.darling | 2 (0.5%) | 193 (0.9%) | 96.5 |
Finally interfacing multivariate traits, although they are now generally integrated out.
120 lines of code changed in 10 files:
Implemented a scaled mixture of Wishart distribution for precision matrix inference.
78 lines of code changed in 1 file:
Allopolyploid model. Added diploidRootIsRoot flag (but only tested code when true), changed formula for network prior, added error check relating to oneHybridization flag, improved debug output.
AlloppChangeNumHybridizations: added error check relating to oneHybridization flag.
AlloppChangeNumHybridizationsParser: added error relating to oneHybridization flag.
AlloppDiploidHistory.java: nicer debug output, changes for diploidRootIsRoot flag.
AlloppLeggedTree: simpler debug output.
AlloppMulLabTree: nicer debug output.
AlloppNetworkNodeSlide: Changed operateOneNodeInDiploidHistory() for diploidRootIsRoot flag.
AlloppNetworkPrior: Changed formula for prior (9.0 -> 7.0*sqrt(#diploids))
AlloppSpeciesBindings: nicer debug output.
AlloppSpeciesNetworkModel: changes for diploidRootIsRoot flag, debug output.
AlloppSpeciesNetworkModelParser: changes for diploidRootIsRoot flag.
68 lines of code changed in 6 files:
Added unit-IntervalLatentLiabilityLikelihood for VNTRs; plan to refactor to remove duplicated code once more fully tested.
8 lines of code changed in 1 file:
Fixed bug in PrecisionMatrixGibbsOperator under power posterior
1 lines of code changed in 1 file:
Updated PrecisionMatrixGibbsOperator for functionality within path sampling from posterior to prior. Should still cause errors in path sampling between two arbitrary models.
13 lines of code changed in 1 file:
Added missing multivariate distribution likelihood model; opens up so many possibilities for rate models. Caution: may have broken Brownian diffusion precision Gibbs sampler.
119 lines of code changed in 1 file:
Allopolyploid model. Added MCMC operator to flip legs and sequences of tetraploid tree to improve mixing. Improved sampling for initial state of network. Added statistic for number of hybridizations.
221 lines of code changed in 3 files:
Allopolyploid model: new version of ChangeNumHybridizations move and bug fixes.
84 lines of code changed in 3 files:
Allopolyploid model: adding code for arbitrary numbers of diploids, tetraploids, hybridizations.
374 lines of code changed in 5 files:
Code for n diploids, one hybridization now working. Experimental code (PopsIO) added for multispecies coalescent with population size parameters integrated out.
100 lines of code changed in 3 files:
Extending allopolyploid model to deal with arbitrary numbers of diploids. Most new code is in new class AlloppDiploidHistory and old class AlloppNetworkNodeSlide.
162 lines of code changed in 1 file:
Improved MCMC moves for allopolyploid model (diploid root, moves of legs).
27 lines of code changed in 1 file:
Adding multiply-labelled tree model. MulSpeciesTreeModel, MulMSCoalescent, MulSpeciesBindings, MulSpeciesTreePrior. Operators MulTreeNodeSlide, MulTreeSequenceReassignment. Plus parsers.
Minor updates to AlloppSpeciesBindings, AlloppSpeciesNetworkModel, the latter dues to changes in logging behaviour.
281 lines of code changed in 2 files:
Loggers weren't logging if they overrode the base class log(int state) - because it had been changed to log(long state). Put a final int version in the base class to prevent overriding.
4 lines of code changed in 2 files:
Enabled Gibbs sampling of precision matrix in NonPhylogeneticMultivariateTraitLikelihood
5 lines of code changed in 1 file:
Fixed integer overflow (when prior sampling) and mixed-math in RandomWalkOperator
7 lines of code changed in 1 file:
Removed some conflicts in XML parsers
0 lines of code changed in 1 file:
Allopolyploid model: Made code compatible with Java 1.5 (removed @Override's). Added citation. More sensible implementation of debugging logger.
0 lines of code changed in 3 files:
438 lines of code changed in 3 files:
Latent liability model appears to work well.
362 lines of code changed in 1 file:
uncommitting an erroneous revision
0 lines of code changed in 1 file:
Trying to get Evolutionary Cartography working...
2 lines of code changed in 1 file:
automatic intelliJ minor syntax fixes
1 lines of code changed in 2 files:
After some more extensive testing, I have now removed the node heights bound check as default. Certain moves still may make this check.
7 lines of code changed in 2 files:
Decoupled the process of checking node heights are within bounds to confirm that an operator is correct from using it to reject a move. By default tree moves just call treeModel.endTreeEdit() to finish editing the tree. If they need to check whether the resulting tree is valid they can call checkTreeIsValid() which throws an exception if not. Most moves do not need this check if correctly implemented. Some I wasn't too sure about so I left the check in for those to give the same behaviour (but it is up to the operator whether this check is made and what to do about it).
TreeModel.endTreeEdit() can optionally make this test (switch on a flag at the top of TreeModel) but throws a runtimeException so stops the program if it fails.
48 lines of code changed in 25 files:
Whilst I test the consequence of removing the height bound checking in TreeModel.endTreeEdit(), I have reinstated the test but it now throws a RuntimeException to see if any operators rely on this test failing to check for invalid moves.
2 lines of code changed in 2 files:
Trunk: @Override is JDK1.6 code.
4 lines of code changed in 2 files:
Adding a new model of hidden linkage among reads in a metagenomic dataset. The model and its application to metagenomic datasets is described in a manuscript in preparation by Aaron Darling and Jonathan Eisen. This is commit phase 1: adding new files. Phase 2 comes next, committing related changes to existing files.
193 lines of code changed in 2 files:
Forgot to actually implement the swap.
4 lines of code changed in 2 files:
Implemented an operator to swap tip trait values in an IntegratedMultivariateTraitLikelihood (yes Marc, I do need to do this).
113 lines of code changed in 1 file:
Add several wrappers to convert automatically between different reductions of TreeTraits (sum over tree, sum across sites, etc.). One can even double-wrap to get sum over tree + sum across sites. This should reduce substantial code-duplication and earn me a gold star.
50 lines of code changed in 2 files:
Porting TipStateSwapOperator over to BEAGLE against my better judgement. Random permutations of tip traits form very poor statistical tests.
66 lines of code changed in 2 files:
Placeholders for a new transition kernel to hopefully solve the uniform internal node prior issues
115 lines of code changed in 1 file:
Placeholders for an awesome new transition kernel to hopefully solve the uniform internal node prior issues
125 lines of code changed in 1 file:
Updating geo-spatial regions for Gibbs sampling
15 lines of code changed in 1 file:
Fixed degrees-of-freedom bug in PrecisionGibbsOperator
11 lines of code changed in 1 file:
Making progress on Gibbs sampling the fully conjugate diffusion model; taking days longer than I expected
29 lines of code changed in 1 file:
Some thoughts on Gibbs sampling the precision matrix
51 lines of code changed in 1 file:
Delegating funcationality in IntegratedMultivariateTraitLikelihood for profiling and future refactoring into a TraitLikelihoodCore
35 lines of code changed in 2 files:
(110 more)