Lines of Code
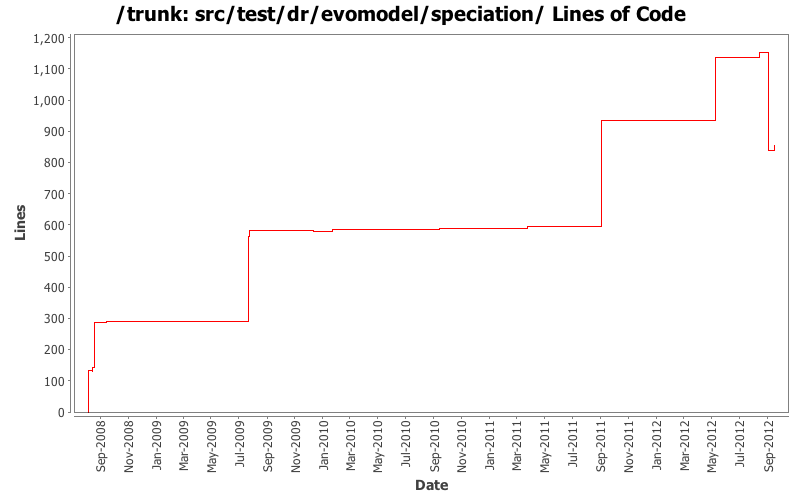
[root]/src/test/dr/evomodel/speciation

| Author | Changes | Lines of Code | Lines per Change |
|---|---|---|---|
| Totals | 63 (100.0%) | 1429 (100.0%) | 22.6 |
| graham853@gmail.com | 6 (9.5%) | 701 (49.1%) | 116.8 |
| alexei.drummond | 18 (28.6%) | 612 (42.8%) | 34.0 |
| dong.w.xie | 21 (33.3%) | 50 (3.5%) | 2.3 |
| dong.w.xie@gmail.com | 3 (4.8%) | 28 (2.0%) | 9.3 |
| jheled | 9 (14.3%) | 26 (1.8%) | 2.8 |
| rambaut | 5 (7.9%) | 11 (0.8%) | 2.2 |
| msuchard | 1 (1.6%) | 1 (0.1%) | 1.0 |
Allopolyploid model: new version of ChangeNumHybridizations move and bug fixes.
18 lines of code changed in 2 files:
Allopolyploid model: adding code for arbitrary numbers of diploids, tetraploids, hybridizations.
100 lines of code changed in 1 file:
trunk: merge Issue 656 : fix p=0 in Birth Death Serial Sampling Model, and update Unit tests.
27 lines of code changed in 2 files:
Code for n diploids, one hybridization now working. Experimental code (PopsIO) added for multispecies coalescent with population size parameters integrated out.
10 lines of code changed in 1 file:
Extending allopolyploid model to deal with arbitrary numbers of diploids. Most new code is in new class AlloppDiploidHistory and old class AlloppNetworkNodeSlide.
234 lines of code changed in 1 file:
trunk: fix Junit test
1 lines of code changed in 1 file:
339 lines of code changed in 1 file:
TRUNK: fix Hudson due to change of BDSS
3 lines of code changed in 1 file:
removed final time interval
1 lines of code changed in 1 file:
Trunk: start to implement Brith Death Serial Sampling in BEAUti.
11 lines of code changed in 1 file:
Added the sampling of the origin of the infection to the birth-death serial sampling model.
1 lines of code changed in 1 file:
Trunk: new change of BirthDeathSerialSamplingModel
2 lines of code changed in 1 file:
Trunk: correct new update of BirthDeathSerialSamplingModel
2 lines of code changed in 1 file:
Trunk: update BirthDeathSerialSamplingModel logL
1 lines of code changed in 1 file:
TRUNK: fix breaken code in test, but wait Alexei to verify the test.
5 lines of code changed in 1 file:
Fixed BirthDeathSerialSampling model. It turns out that death needs to be parameterized relative to birth to get good mixing as they are extremely highly correlated in the data sets I have looked at. I have added the option to parameterize relative or absolute death rate.
1 lines of code changed in 1 file:
Trunk refactoring: organize tree and treelikelihood parsers.
6 lines of code changed in 7 files:
Trunk refactoring: finish coalescent (split parsers).
4 lines of code changed in 2 files:
Trunk: hide the confused constructor from MCMC.
2 lines of code changed in 2 files:
Trunk: fix seed for integration test, and see whether to solve unstable Hudson test.
10 lines of code changed in 2 files:
Failed to update a few files after last commit..
6 lines of code changed in 2 files:
Replaced 'preBurnin' with an autoOptimizeDelay (coercionDelay). The rationale for this is that the preBurnin meant that the initial state was not in the log files, it was unecessary and it cause a small inefficiency as the JIT compiler re-optimized after the preburnin chain had finished. We also didn't see warning messages until after the pre-burnin period. There is a new attribute to the MCMC parser - autoOptimizeDelay (to which the old preBurnin attribute now maps so old XML will run).
2 lines of code changed in 2 files:
tidy
3 lines of code changed in 1 file:
fix test failure
2 lines of code changed in 2 files:
Add types of trees to birth-death density
8 lines of code changed in 3 files:
share constants via variables
11 lines of code changed in 1 file:
fixed test...
8 lines of code changed in 2 files:
birth-death-serial-sampling-model testing... appears that birth-death-gernhard-08 gives wrong likelihoods. (see BirthDeathLikelihoodTest)
16 lines of code changed in 2 files:
Tanja's Birth-Death-Serial-Sampling tree prior. Doesn't quite match up with BirthDeathGernhard08 likelihoods yet...
272 lines of code changed in 3 files:
Getting linear model sampling with BSSVS working.
1 lines of code changed in 1 file:
A small improvment in reducing usage of litteral string constants.
Each constant should be defined in one place and referenced elsewhere.
The best place I found is in the various parsers. This may be wrong but now moving it elsewhere is a simple matter.
2 lines of code changed in 1 file:
Correct the committed JUnit test files and make them working.
4 lines of code changed in 2 files:
Sampling proportion added to Birth-Death, based on Tanja derivation. Not tested by simulation. Expressions are identical to the ones pre-change when the default proportion (1) is used.
1 lines of code changed in 1 file:
Remove depracated use of constant demographic in Wilson Balding operator
Merge some uses of the "rootHeight" constant string
2 lines of code changed in 1 file:
Added the sequencing error/DNA damage model options to BEAUti.
3 lines of code changed in 1 file:
tidying operators
1 lines of code changed in 1 file:
refactored random local stuff so that it can be used for both clock rate and speciation models
1 lines of code changed in 1 file:
added special logger for random local yule model
1 lines of code changed in 1 file:
Added RandomLocalYuleModel and a new move for stochastical variable selection models on trees
143 lines of code changed in 1 file:
added new tests for wide exchange operator in ExchangeOperatorTest and YuleModelTest... I would encourage other BEAST developers to consider writing JUnit tests... they are easy and fun!
23 lines of code changed in 1 file:
(2 more)